Image Analysis
SCI's imaging work addresses fundamental questions in 2D and 3D image processing, including filtering, segmentation, surface reconstruction, and shape analysis. In low-level image processing, this effort has produce new nonparametric methods for modeling image statistics, which have resulted in better algorithms for denoising and reconstruction. Work with particle systems has led to new methods for visualizing and analyzing 3D surfaces. Our work in image processing also includes applications of advanced computing to 3D images, which has resulted in new parallel algorithms and real-time implementations on graphics processing units (GPUs). Application areas include medical image analysis, biological image processing, defense, environmental monitoring, and oil and gas.
Ross Whitaker
Segmentation
Chris Johnson
Diffusion Tensor AnalysisFunded Research Projects:
Publications in Image Analysis:
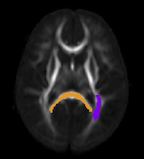
  Altered corpus callosum morphology associated with autism over the first 2 years of life J. J. Wolff, G. Gerig, J. D. Lewis, T. Soda, M. A. Styner, C. Vachet, K. N. Botteron, J. T. Elison, S. R. Dager, A. M. Estes, H. C. Hazlett, R. T. Schultz, L. Zwaigenbaum, J. Piven. In Brain, 2015. DOI: 10.1093/brain/awv118 Numerous brain imaging studies indicate that the corpus callosum is smaller in older children and adults with autism spectrum disorder. However, there are no published studies examining the morphological development of this connective pathway in infants at-risk for the disorder. Magnetic resonance imaging data were collected from 270 infants at high familial risk for autism spectrum disorder and 108 low-risk controls at 6, 12 and 24 months of age, with 83% of infants contributing two or more data points. Fifty-seven children met criteria for ASD based on clinical-best estimate diagnosis at age 2 years. Corpora callosa were measured for area, length and thickness by automated segmentation. We found significantly increased corpus callosum area and thickness in children with autism spectrum disorder starting at 6 months of age. These differences were particularly robust in the anterior corpus callosum at the 6 and 12 month time points. Regression analysis indicated that radial diffusivity in this region, measured by diffusion tensor imaging, inversely predicted thickness. Measures of area and thickness in the first year of life were correlated with repetitive behaviours at age 2 years. In contrast to work from older children and adults, our findings suggest that the corpus callosum may be larger in infants who go on to develop autism spectrum disorder. This result was apparent with or without adjustment for total brain volume. Although we did not see a significant interaction between group and age, cross-sectional data indicated that area and thickness differences diminish by age 2 years. Regression data incorporating diffusion tensor imaging suggest that microstructural properties of callosal white matter, which includes myelination and axon composition, may explain group differences in morphology. |
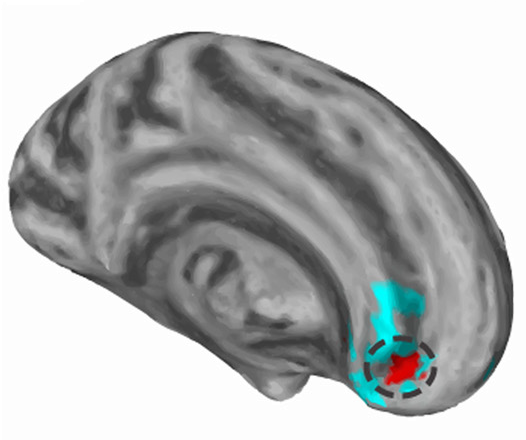
  Prenatal Drug Exposure Affects Neonatal Brain Functional Connectivity A. P. Salzwedel, K. M. Grewen, C. Vachet, G. Gerig, W. Lin,, W. Gao. In The Journal of Neuroscience, Vol. 35, No. 14, pp. 5860-5869. April, 2015. DOI: 10.1523/JNEUROSCI.4333-14.2015 Prenatal drug exposure, particularly prenatal cocaine exposure (PCE), incurs great public and scientific interest because of its associated neurodevelopmental consequences. However, the neural underpinnings of PCE remain essentially uncharted, and existing studies in school-aged children and adolescents are confounded greatly by postnatal environmental factors. In this study, leveraging a large neonate sample (N = 152) and non-invasive resting-state functional magnetic resonance imaging, we compared human infants with PCE comorbid with other drugs (such as nicotine, alcohol, marijuana, and antidepressant) with infants with similar non-cocaine poly drug exposure and drug-free controls. We aimed to characterize the neural correlates of PCE based on functional connectivity measurements of the amygdala and insula at the earliest stage of development. Our results revealed common drug exposure-related connectivity disruptions within the amygdala–frontal, insula–frontal, and insula–sensorimotor circuits. Moreover, a cocaine-specific effect was detected within a subregion of the amygdala–frontal network. This pathway is thought to play an important role in arousal regulation, which has been shown to be irregular in PCE infants and adolescents. These novel results provide the earliest human-based functional delineations of the neural-developmental consequences of prenatal drug exposure and thus open a new window for the advancement of effective strategies aimed at early risk identification and intervention. |
 Spatio-temporal Image Analysis for Longitudinal and Time-Series Image Data, S. Durrleman, T.P. Fletcher, G. Gerig, M. Niethammer, X. Pennec (Eds.). In Proceedings of the Third International Workshop, STIA 2014, Image Processing, Computer Vision, Pattern Recognition, and Graphics, Vol. 8682, Springer LNCS, 2015. ISBN: 978-3-319-14905-9 This book constitutes the thoroughly refereed post-conference proceedings of the Third |
  Cell tracking using particle filters with implicit convex shape model in 4D confocal microscopy images N. Ramesh, T. Tasdizen. In 2014 IEEE International Conference on Image Processing (ICIP), IEEE, Oct, 2014. DOI: 10.1109/icip.2014.7025089 Bayesian frameworks are commonly used in tracking algorithms. An important example is the particle filter, where a stochastic motion model describes the evolution of the state, and the observation model relates the noisy measurements to the state. Particle filters have been used to track the lineage of cells. Propagating the shape model of the cell through the particle filter is beneficial for tracking. We approximate arbitrary shapes of cells with a novel implicit convex function. The importance sampling step of the particle filter is defined using the cost associated with fitting our implicit convex shape model to the observations. Our technique is capable of tracking the lineage of cells for nonmitotic stages. We validate our algorithm by tracking the lineage of retinal and lens cells in zebrafish embryos. |
 Multiatlas Segmentation as Nonparametric Regression S.P. Awate, R.T. Whitaker. In IEEE Trans Med Imaging, April, 2014. PubMed ID: 24802528 This paper proposes a novel theoretical framework to model and analyze the statistical characteristics of a wide range of segmentation methods that incorporate a database of label maps or atlases; such methods are termed as label fusion or multiatlas segmentation. We model these multiatlas segmentation problems as nonparametric regression problems in the high-dimensional space of image patches. We analyze the nonparametric estimator's convergence behavior that characterizes expected segmentation error as a function of the size of the multiatlas database. We show that this error has an analytic form involving several parameters that are fundamental to the specific segmentation problem (determined by the chosen anatomical structure, imaging modality, registration algorithm, and labelfusion algorithm). We describe how to estimate these parameters and show that several human anatomical structures exhibit the trends modeled analytically. We use these parameter estimates to optimize the regression estimator. We show that the expected error for large database sizes is well predicted by models learned on small databases. Thus, a few expert segmentations can help predict the database sizes required to keep the expected error below a specified tolerance level. Such cost-benefit analysis is crucial for deploying clinical multiatlas segmentation systems. |
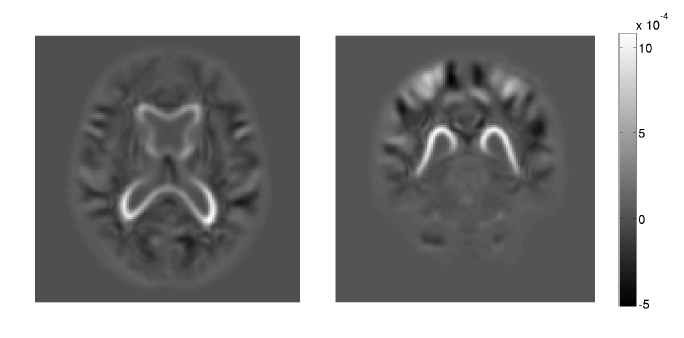
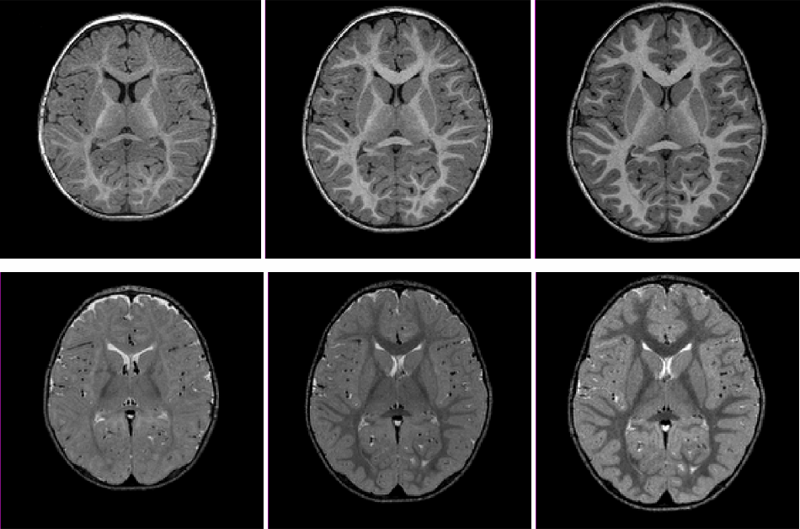
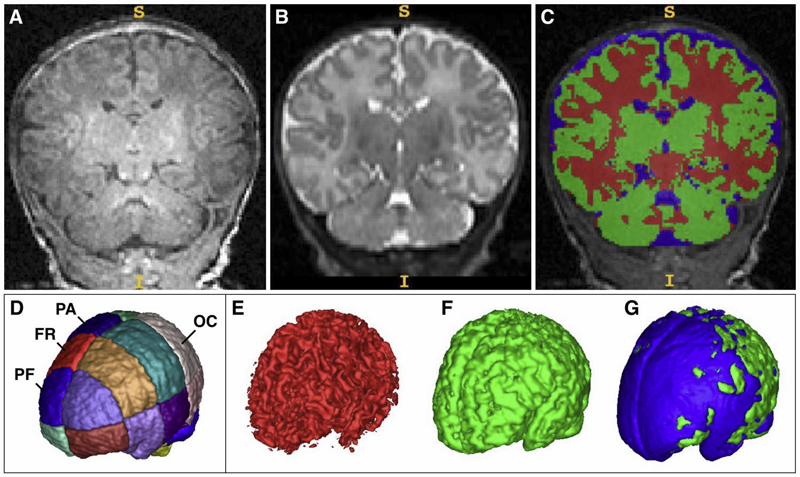
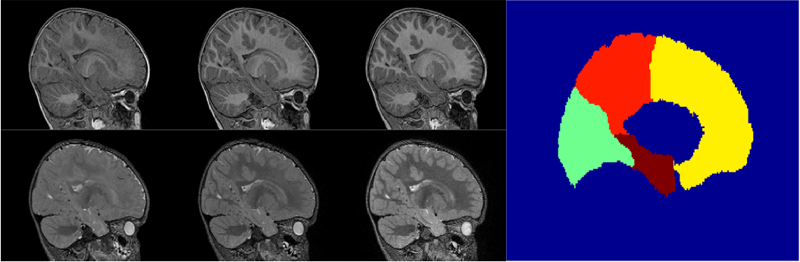
  Prenatal cocaine effects on brain structure in early infancy K. Grewen, M. Burchinal, C. Vachet, S. Gouttard, J.H. Gilmore, W. Lin, J. Johns, M. Elam, G. Gerig. In NeuroImage, Vol. 101, pp. 114--123. November, 2014. DOI: 10.1016/j.neuroimage.2014.06.070 Prenatal cocaine exposure (PCE) is related to subtle deficits in cognitive and behavioral function in infancy, childhood and adolescence. Very little is known about the effects of in utero PCE on early brain development that may contribute to these impairments. The purpose of this study was to examine brain structural differences in infants with and without PCE. We conducted MRI scans of newborns (mean age = 5 weeks) to determine cocaine's impact on early brain structural development. Subjects were three groups of infants: 33 with PCE co-morbid with other drugs, 46 drug-free controls and 40 with prenatal exposure to other drugs (nicotine, alcohol, marijuana, opiates, SSRIs) but without cocaine. Infants with PCE exhibited lesser total gray matter (GM) volume and greater total cerebral spinal fluid (CSF) volume compared with controls and infants with non-cocaine drug exposure. Analysis of regional volumes revealed that whole brain GM differences were driven primarily by lesser GM in prefrontal and frontal brain regions in infants with PCE, while more posterior regions (parietal, occipital) did not differ across groups. Greater CSF volumes in PCE infants were present in prefrontal, frontal and parietal but not occipital regions. Greatest differences (GM reduction, CSF enlargement) in PCE infants were observed in dorsal prefrontal cortex. Results suggest that PCE is associated with structural deficits in neonatal cortical gray matter, specifically in prefrontal and frontal regions involved in executive function and inhibitory control. Longitudinal study is required to determine whether these early differences persist and contribute to deficits in cognitive functions and enhanced risk for drug abuse seen at school age and in later life. |
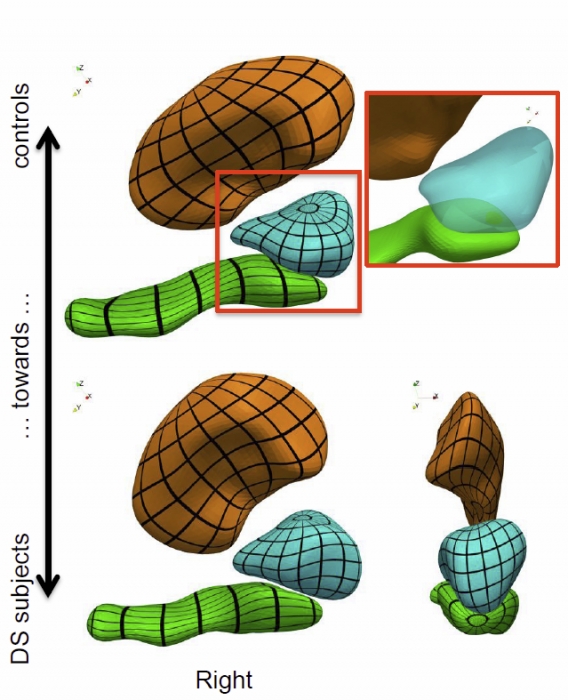
  Morphometry of anatomical shape complexes with dense deformations and sparse parameters S. Durrleman, M. Prastawa, N. Charon, J.R. Korenberg, S. Joshi, G. Gerig, A. Trouvé. In NeuroImage, 2014. DOI: 10.1016/j.neuroimage.2014.06.043 We propose a generic method for the statistical analysis of collections of anatomical shape complexes, namely sets of surfaces that were previously segmented and labeled in a group of subjects. The method estimates an anatomical model, the template complex, that is representative of the population under study. Its shape reflects anatomical invariants within the dataset. In addition, the method automatically places control points near the most variable parts of the template complex. Vectors attached to these points are parameters of deformations of the ambient 3D space. These deformations warp the template to each subject’s complex in a way that preserves the organization of the anatomical structures. Multivariate statistical analysis is applied to these deformation parameters to test for group differences. Results of the statistical analysis are then expressed in terms of deformation patterns of the template complex, and can be visualized and interpreted. The user needs only to specify the topology of the template complex and the number of control points. The method then automatically estimates the shape of the template complex, the optimal position of control points and deformation parameters. The proposed approach is completely generic with respect to any type of application and well adapted to efficient use in clinical studies, in that it does not require point correspondence across surfaces and is robust to mesh imperfections such as holes, spikes, inconsistent orientation or irregular meshing. The approach is illustrated with a neuroimaging study of Down syndrome (DS). Results demonstrate that the complex of deep brain structures shows a statistically significant shape difference between control and DS subjects. The deformation-based modelingis able to classify subjects with very high specificity and sensitivity, thus showing important generalization capability even given a low sample size. We show that results remain significant even if the number of control points, and hence the dimension of variables in the statistical model, are drastically reduced. The analysis may even suggest that parsimonious models have an increased statistical performance. The method has been implemented in the software Deformetrica, which is publicly available at www.deformetrica.org.
Keywords: morphometry, deformation, varifold, anatomy, shape, statistics |
  GuideME: Slice-guided Semiautomatic Multivariate Exploration of Volumes L. Zhou, C.D. Hansen. In Computer Graphics Forum, Vol. 33, No. 3, Wiley-Blackwell, pp. 151--160. jun, 2014. DOI: 10.1111/cgf.12371 Multivariate volume visualization is important for many applications including petroleum exploration and medicine. State-of-the-art tools allow users to interactively explore volumes with multiple linked parameter-space views. However, interactions in the parameter space using trial-and-error may be unintuitive and time consuming. Furthermore, switching between different views may be distracting. In this paper, we propose GuideME: a novel slice-guided semiautomatic multivariate volume exploration approach. Specifically, the approach comprises four stages: attribute inspection, guided uncertainty-aware lasso creation, automated feature extraction and optional spatial fine tuning and visualization. Throughout the exploration process, the user does not need to interact with the parameter views at all and examples of complex real-world data demonstrate the usefulness, efficiency and ease-of-use of our method. |
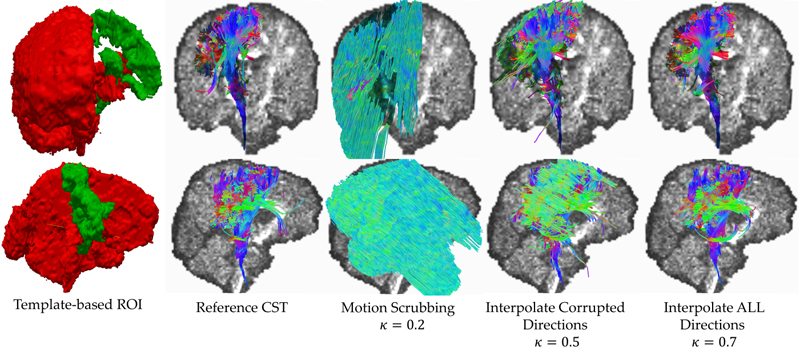
  Network inefficiencies in autism spectrum disorder at 24 months J.D. Lewis, A.C. Evans, J.R. Pruett, K. Botteron, L. Zwaigenbaum, A. Estes, G. Gerig, L. Collins, P. Kostopoulos, R. McKinstry, S. Dager, S. Paterson, R. Schultz, M. Styner, H. Hazlett, J. Piven, IBIS network. In Translational Psychiatry, Vol. 4, No. 5, Nature Publishing Group, pp. e388. May, 2014. DOI: 10.1038/tp.2014.24 Autism Spectrum Disorder (ASD) is a developmental disorder defined by behavioural symptoms that emerge during the first years of life. Associated with these symptoms are differences in the structure of a wide array of brain regions, and in the connectivity between these regions. However, the use of cohorts with large age variability and participants past the generally recognized age of onset of the defining behaviours means that many of the reported abnormalities may be a result of cascade effects of developmentally earlier deviations. This study assessed differences in connectivity in ASD at the age at which the defining behaviours first become clear. The participants were 113 24-month-olds at high risk for ASD, 31 of whom were classified as ASD, and 23 typically developing 24-month-olds at low risk for ASD. Utilizing diffusion data to obtain measures of the length and strength of connections between anatomical regions, we performed an analysis of network efficiency. Our results showed significantly decreased local and global efficiency over temporal, parietal, and occipital lobes in high-risk infants classified as ASD, relative to both low- and high-risk infants not classified as ASD. The frontal lobes showed only a reduction in global efficiency in Broca's area. Additionally, these same regions showed an inverse relation between efficiency and symptom severity across the high-risk infants. The results suggest delay or deficits in infants with ASD in the optimization of both local and global aspects of network structure in regions involved in processing auditory and visual stimuli, language, and nonlinguistic social stimuli. Keywords: autism, infant siblings, connectivity, network analysis, efficiency |