To download installation packages for Windows/Mac/Linux and/or the source code, please visit https://github.com/SCIInstitute/ShapeWorks/releases/tag/v6.3.0
What is new in ShapeWorks 6.3?
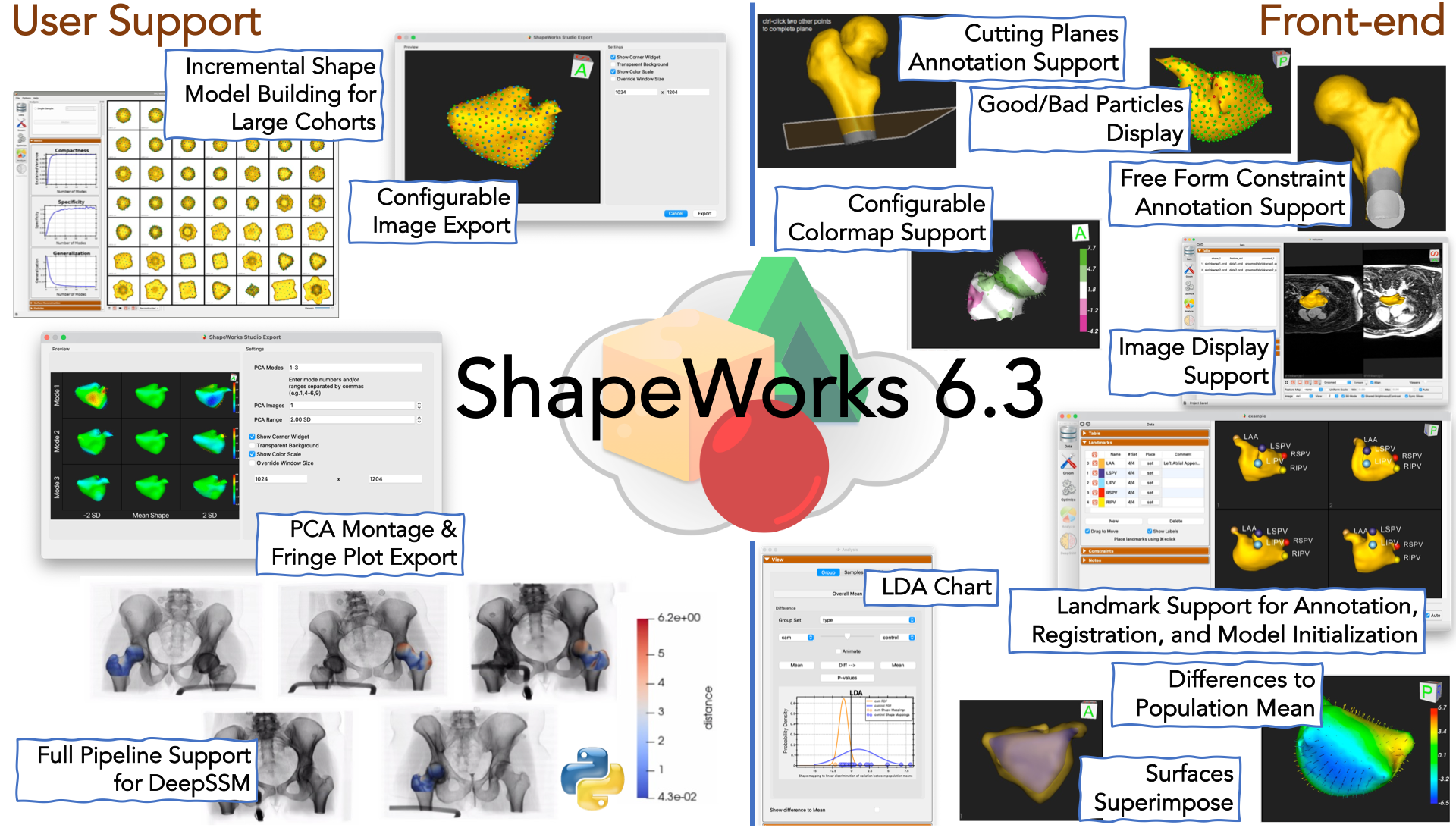
New Features in Studio
- Support for landmark definitions, alignments using landmarks and initial particles from landmarks. For more details, see the Studio Data Module Documentation
- UI for defining cutting planes and free form constraints. For more details, see the Studio Data Module Documentation
- Image volume support: New support has been added for displaying 2D slices from image volumes (e.g. CT/MRI). For more details, see Image Volume Support
- New selectable and configurable colormaps
- Added new support for showing the difference to the mean for any given mesh (subjects or generated PCA mode positions). See Difference to Mean.
- Added new support for displaying multiple image types (e.g. original vs groomed) with individual opacity settings. Also ability to show surface to surface distance. See Comparing mesh types.
- A new image export dialog as been added with various export options. See Image Export
- PCA Montage and Fringe plot export. Building on the image export dialog, the PCA Montage exporter allows you to create a multi-image montage across PCA modes. See PCA montage.
- Export scalar values: Addition export options have been added to export mesh scalars, particle scalars, and all subjects particle scalars.
- Procrustes scaling only mode: New support for running procrustes in a scaling-only mode has been added.
- Good/bad particle display: The Particles Panel enabled the display of "good/bad particles" in ShapeWorks Studio.
- Group LDA chart in Studio: Support for the group LDA chart has been added in Studio
Improved Use Cases and Documentation
- Added grooming steps to mesh-based use cases using the mesh Python API
- Alignment transforms are now passed to the optimizer and used in optimization instead of being applied before optimization. This results in local particles in the original data's coordinate system, allowing for easier subsequent analysis
- The use cases now use project spreadsheets in optimizations instead of XML files. This format is more interpretable and allows of better integration with Studio. The project sheets support multiple domains, fixed domains, constraints
- The femur use case has been refactored into a single use case where alignment transforms and cutting plane constraints are passed in optimization.
- Grooming added for multiple domain use cases. The pipeline demonstrates alignment w.r.t domain 1 ellipsoids.
- Added a Studio use case for constraints and a pseudo-tutorial for it in the documentation.
DeepSSM Use Case:
- The DeepSSM use case has been updated to demonstrate the full pipeline, including training data generation instead of relying on the femur use case to create a training shape model.
- The use case now demonstrates how to optimize validation particles via fixed domain optimization where the training particles are unchanged.
- Image-to-image registration tools have been added to prepare DeepSSM input images without requiring corresponding segmentations or meshes. This allows for true inference with DeepSSM.
- A new use case has been added, demonstrating how a shape model can be optimized incrementally on 3D supershapes. This approach is beneficial when the cohort of shapes is very large, and single optimization would be slow, and when the dataset is small but contains a large amount of shape variation.
- Functionality has been added to select the order of shape optimization based on the distance of each shape to all others in the cohort. This allows for particles to be fit to inlier shapes first, then outliers.
- Documentation has been added that explains the use case and quantitatively demonstrates the benefit of incremental optimization.
ShapeWorks Back-end
- Added constraints functionality for the mesh domain both clipping and augmented lagrangian together with a flag to flip between the two options.
- Group Difference Statistics in Python can now perform LDA. The use case also demonstrates Linear Discrimination of Variation (LDA) for analyzing shape variation between the subgroups.
Fixes
- Studio: TabWidget rendering on MacOS 11/12 fixed
- Mesh::toDistanceTransform fixed
- Studio: Fixed optimization abort not always aborting
- Optimize: Fixed particle splitting for use with input transforms
- Studio: Fix clamping of glyph size
- Studio: Fix bug when groom output path is blank
