SCI Publications
2014
A. J. Perez, M. Seyedhosseini, T. J. Deerinck, E. A. Bushong, S. Panda, T. Tasdizen, M. H. Ellisman.
“A workflow for the automatic segmentation of organelles in electron microscopy image stacks,” In Frontiers in Neuroanatomy, Vol. 8, No. 126, 2014.
DOI: 10.3389/fnana.2014.00126
Electron microscopy (EM) facilitates analysis of the form, distribution, and functional status of key organelle systems in various pathological processes, including those associated with neurodegenerative disease. Such EM data often provide important new insights into the underlying disease mechanisms. The development of more accurate and efficient methods to quantify changes in subcellular microanatomy has already proven key to understanding the pathogenesis of Parkinson's and Alzheimer's diseases, as well as glaucoma. While our ability to acquire large volumes of 3D EM data is progressing rapidly, more advanced analysis tools are needed to assist in measuring precise three-dimensional morphologies of organelles within data sets that can include hundreds to thousands of whole cells. Although new imaging instrument throughputs can exceed teravoxels of data per day, image segmentation and analysis remain significant bottlenecks to achieving quantitative descriptions of whole cell structural organellomes. Here, we present a novel method for the automatic segmentation of organelles in 3D EM image stacks. Segmentations are generated using only 2D image information, making the method suitable for anisotropic imaging techniques such as serial block-face scanning electron microscopy (SBEM). Additionally, no assumptions about 3D organelle morphology are made, ensuring the method can be easily expanded to any number of structurally and functionally diverse organelles. Following the presentation of our algorithm, we validate its performance by assessing the segmentation accuracy of different organelle targets in an example SBEM dataset and demonstrate that it can be efficiently parallelized on supercomputing resources, resulting in a dramatic reduction in runtime.
A. Perez, M. Seyedhosseini, T. Tasdizen, M. Ellisman.
“Automated workflows for the morphological characterization of organelles in electron microscopy image stacks (LB72),” In The FASEB Journal, Vol. 28, No. 1 Supplement LB72, April, 2014.
Advances in three-dimensional electron microscopy (EM) have facilitated the collection of image stacks with a field-of-view that is large enough to cover a significant percentage of anatomical subdivisions at nano-resolution. When coupled with enhanced staining protocols, such techniques produce data that can be mined to establish the morphologies of all organelles across hundreds of whole cells in their in situ environments. Although instrument throughputs are approaching terabytes of data per day, image segmentation and analysis remain significant bottlenecks in achieving quantitative descriptions of whole cell organellomes. Here we describe computational workflows that achieve the automatic segmentation of organelles from regions of the central nervous system by applying supervised machine learning algorithms to slices of serial block-face scanning EM (SBEM) datasets. We also demonstrate that our workflows can be parallelized on supercomputing resources, resulting in a dramatic reduction of their run times. These methods significantly expedite the development of anatomical models at the subcellular scale and facilitate the study of how these models may be perturbed following pathological insults.
N. Ramesh, T. Tasdizen.
“Cell tracking using particle filters with implicit convex shape model in 4D confocal microscopy images,” In 2014 IEEE International Conference on Image Processing (ICIP), IEEE, Oct, 2014.
DOI: 10.1109/icip.2014.7025089
Bayesian frameworks are commonly used in tracking algorithms. An important example is the particle filter, where a stochastic motion model describes the evolution of the state, and the observation model relates the noisy measurements to the state. Particle filters have been used to track the lineage of cells. Propagating the shape model of the cell through the particle filter is beneficial for tracking. We approximate arbitrary shapes of cells with a novel implicit convex function. The importance sampling step of the particle filter is defined using the cost associated with fitting our implicit convex shape model to the observations. Our technique is capable of tracking the lineage of cells for nonmitotic stages. We validate our algorithm by tracking the lineage of retinal and lens cells in zebrafish embryos.
F. Rousset, C. Vachet, C. Conlin, M. Heilbrun, J.L. Zhang, V.S. Lee, G. Gerig.
“Semi-automated application for kidney motion correction and filtration analysis in MR renography,” In Proceeding of the 2014 Joint Annual Meeting ISMRM-ESMRMB, pp. (accepted). 2014.
Altered renal function commonly affects patients with cirrhosis, a consequence of chronic liver disease. From lowdose contrast material-enhanced magnetic resonance (MR) renography, we can estimate the Glomerular Filtration Rate (GFR), an important parameter to assess renal function. Two-dimensional MR images are acquired every 2 seconds for approximately 5 minutes during free breathing, which results in a dynamic series of 140 images representing kidney filtration over time. This specific acquisition presents dynamic contrast changes but is also challenged by organ motion due to breathing. Rather than use conventional image registration techniques, we opted for an alternative method based on object detection. We developed a novel analysis framework available under a stand-alone toolkit to efficiently register dynamic kidney series, manually select regions of interest, visualize the concentration curves for these ROIs, and fit them into a model to obtain GFR values. This open-source cross-platform application is written in C++, using the Insight Segmentation and Registration Toolkit (ITK) library, and QT4 as a graphical user interface.
N. Sadeghi, J.H. Gilmore, W. Lin, G. Gerig.
“Normative Modeling of Early Brain Maturation from Longitudinal DTI Reveals Twin-Singleton Differences,” In Proceeding of the 2014 Joint Annual Meeting ISMRM-ESMRMB, pp. (accepted). 2014.
Early brain development of white matter is characterized by rapid organization and structuring. Magnetic Resonance diffusion tensor imaging (MR-DTI) provides the possibility of capturing these changes non-invasively by following individuals longitudinally in order to better understand departures from normal brain development in subjects at risk for mental illness [1]. Longitudinal imaging of individuals suggests the use of 4D (3D, time) image analysis and longitudinal statistical modeling [3].
N. Sadeghi, P.T. Fletcher, M. Prastawa, J.H. Gilmore, G. Gerig.
“Subject-specific prediction using nonlinear population modeling: Application to early brain maturation from DTI,” In Proceedings of Medical Image Computing and Computer-Assisted Intervention (MICCAI 2014), 2014.
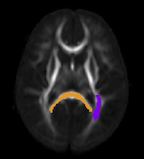
A.R. Sanderson.
“An Alternative Formulation of Lyapunov Exponents for Computing Lagrangian Coherent Structures,” In Proceedings of the 2014 IEEE Pacific Visualization Symposium (PacificVis), Yokahama Japan, 2014.
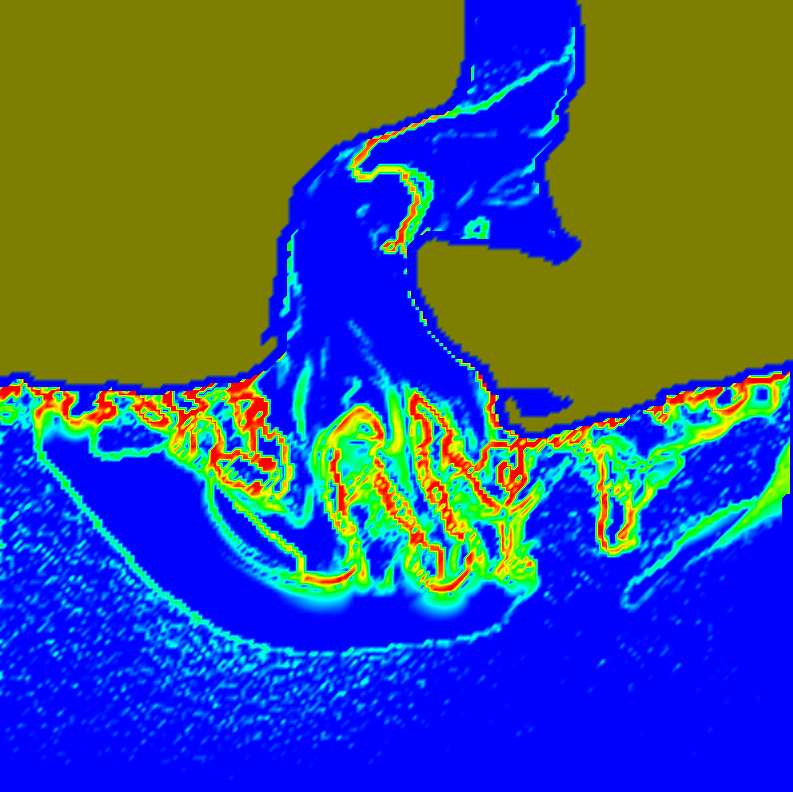
M. Seyedhosseini, T. Tasdizen.
“Disjunctive normal random forests,” In Pattern Recognition, September, 2014.
DOI: 10.1016/j.patcog.2014.08.023
We develop a novel supervised learning/classification method, called disjunctive normal random forest (DNRF). A DNRF is an ensemble of randomly trained disjunctive normal decision trees (DNDT). To construct a DNDT, we formulate each decision tree in the random forest as a disjunction of rules, which are conjunctions of Boolean functions. We then approximate this disjunction of conjunctions with a differentiable function and approach the learning process as a risk minimization problem that incorporates the classification error into a single global objective function. The minimization problem is solved using gradient descent. DNRFs are able to learn complex decision boundaries and achieve low generalization error. We present experimental results demonstrating the improved performance of DNDTs and DNRFs over conventional decision trees and random forests. We also show the superior performance of DNRFs over state-of-the-art classification methods on benchmark datasets.
Keywords: Random forest, Decision tree, Classifier, Supervised learning, Disjunctive normal form
M. Seyedhosseini, T. Tasdizen.
“Scene Labeling with Contextual Hierarchical Models,” In CoRR, Vol. abs/1402.0595, 2014.
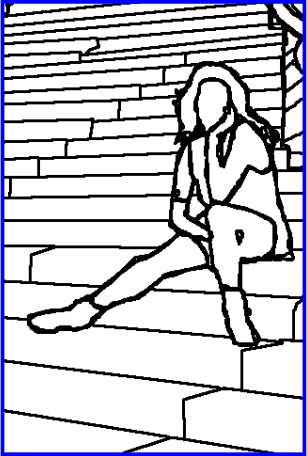

A. Sharma, P.T. Fletcher, J.H. Gilmore, M.L. Escolar, A. Gupta, M. Styner, G. Gerig.
“Parametric Regression Scheme for Distributions: Analysis of DTI Fiber Tract Diffusion Changes in Early Brain Development,” In Proceedings of the 2014 IEEE International Symposium on Biomedical Imaging (ISBI), pp. (accepted). 2014.
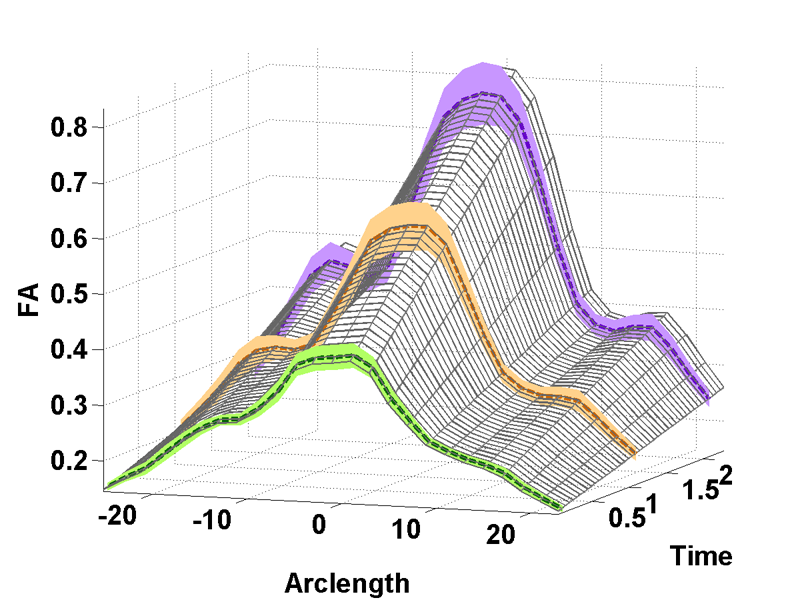
Keywords: linear regression, distribution-valued data, spatiotemporal growth trajectory, DTI, early neurodevelopment
N.P. Singh, J. Hinkle, S. Joshi, P.T. Fletcher.
“An Efficient Parallel Algorithm for Hierarchical Geodesic Models in Diffeomorphisms,” In Proceedings of the 2014 IEEE International Symposium on Biomedical Imaging (ISBI), pp. (accepted). 2014.
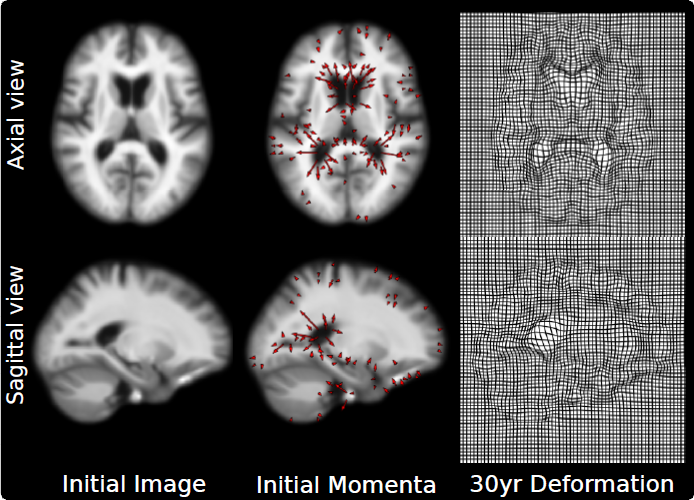
Keywords: LDDMM, HGM, Vector Momentum, Diffeomorphisms, Longitudinal Analysis
P. Skraba, Bei Wang, G. Chen, P. Rosen.
“2D Vector Field Simplification Based on Robustness,” In Proceedings of the 2014 IEEE Pacific Visualization Symposium, PacificVis, Note: Awarded Best Paper!, 2014.
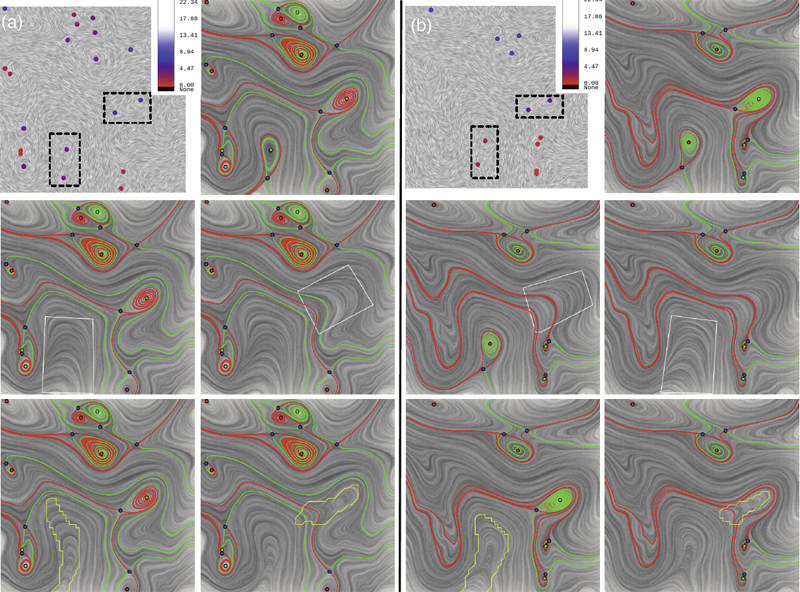
Keywords: vector field, topology-based techniques, flow visualization
P. Skraba, Bei Wang.
“Interpreting Feature Tracking Through the Lens of Robustness,” In Mathematics and Visualization, Springer, pp. 19-37. 2014.
DOI: 10.1007/978-3-319-04099-8_2
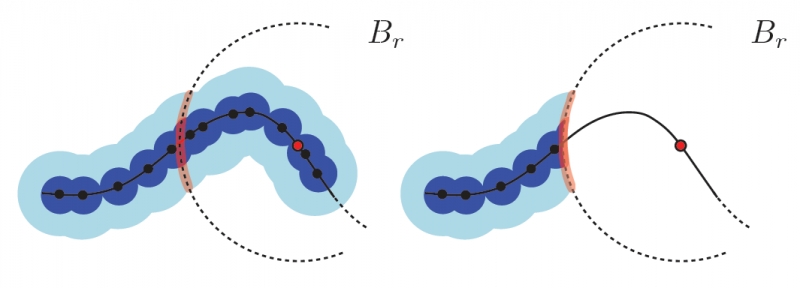
P. Skraba, Bei Wang.
“Approximating Local Homology from Samples,” In Proceedings 25th Annual ACM-SIAM Symposium on Discrete Algorithms (SODA), pp. 174-192. 2014.
R. Stoll, E. Pardyjak, J.J. Kim, T. Harman, A.N. Hayati.
“An inter-model comparison of three computation fluid dynamics techniques for step-up and step-down street canyon flows,” In ASME FEDSM/ICNMM symposium on urban fluid mechanics, August, 2014.
M. Streit, A. Lex, S. Gratzl, C. Partl, D. Schmalstieg, H. Pfister, P. J. Park,, N. Gehlenborg.
“Guided visual exploration of genomic stratifications in cancer,” In Nature Methods, Vol. 11, No. 9, pp. 884--885. Sep, 2014.
ISSN: 1548-7091
DOI: 10.1038/nmeth.3088
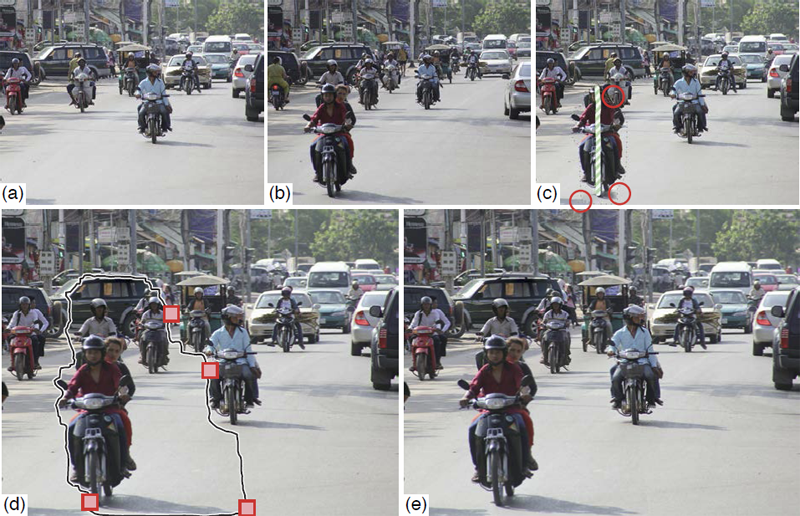
B. Summa, A.A. Gooch, G. Scorzelli, V. Pascucci.
“Towards Paint and Click: Unified Interactions for Image Boundaries,” SCI Technical Report, No. UUSCI-2014-004, SCI Institute, University of Utah, December, 2014.

T. Tasdizen, M. Seyedhosseini, T. Liu, C. Jones, E. Jurrus.
“Image Segmentation for Connectomics Using Machine Learning,” In Computational Intelligence in Biomedical Imaging, Edited by Suzuki, Kenji, Springer New York, pp. 237--278. 2014.
ISBN: 978-1-4614-7244-5
DOI: 10.1007/978-1-4614-7245-2_10
Reconstruction of neural circuits at the microscopic scale of individual neurons and synapses, also known as connectomics, is an important challenge for neuroscience. While an important motivation of connectomics is providing anatomical ground truth for neural circuit models, the ability to decipher neural wiring maps at the individual cell level is also important in studies of many neurodegenerative diseases. Reconstruction of a neural circuit at the individual neuron level requires the use of electron microscopy images due to their extremely high resolution. Computational challenges include pixel-by-pixel annotation of these images into classes such as cell membrane, mitochondria and synaptic vesicles and the segmentation of individual neurons. State-of-the-art image analysis solutions are still far from the accuracy and robustness of human vision and biologists are still limited to studying small neural circuits using mostly manual analysis. In this chapter, we describe our image analysis pipeline that makes use of novel supervised machine learning techniques to tackle this problem.
C. Turkay, A. Lex, M. Streit, H. Pfister,, H. Hauser.
“Characterizing Cancer Subtypes using Dual Analysis in Caleydo,” In IEEE Computer Graphics and Applications, Vol. 34, No. 2, pp. 38--47. March, 2014.
ISSN: 0272-1716
DOI: 10.1109/MCG.2014.1
Dual analysis uses statistics to describe both the dimensions and rows of a high-dimensional dataset. Researchers have integrated it into StratomeX, a Caleydo view for cancer subtype analysis. In addition, significant-difference plots show the elements of a candidate subtype that differ significantly from other subtypes, thus letting analysts characterize subtypes. Analysts can also investigate how data samples relate to their assigned subtype and other groups. This approach lets them create well-defined subtypes based on statistical properties. Three case studies demonstrate the approach's utility, showing how it reproduced findings from a published subtype characterization.
C.J. Underwood, L.T. Edgar, J.B. Hoying, J.A. Weiss.
“Cell-generated traction forces and the resulting matrix deformation modulate microvascular alignment and growth during angiogenesis,” In American Journal of Physiology: Heart and Circulatory Physiology, Vol. 307, No. H152-H164, 2014.
DOI: 10.1152/ajpheart.00995.2013
PubMed ID: 24816262
PubMed Central ID: PMC4101638
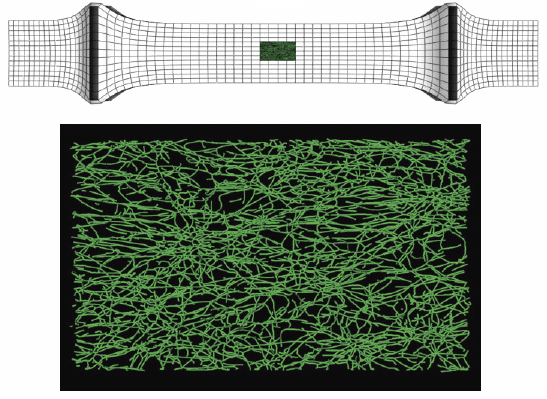

Keywords: angiogenesis, deformation, image analysis, morphometry, orientation, strain
Page 42 of 144
