SCI Publications
2014


S. Gratzl, N. Gehlenborg, A. Lex, H. Pfister, M. Streit.
“Domino: Extracting, Comparing, and Manipulating Subsets across Multiple Tabular Datasets,” In IEEE Transactions on Visualization and Computer Graphics (InfoVis '14), Vol. 20, No. 12, pp. 2023--2032. 2014.
ISSN: 1077-2626
DOI: 10.1109/TVCG.2014.2346260
Answering questions about complex issues often requires analysts to take into account information contained in multiple interconnected datasets. A common strategy in analyzing and visualizing large and heterogeneous data is dividing it into meaningful subsets. Interesting subsets can then be selected and the associated data and the relationships between the subsets visualized. However, neither the extraction and manipulation nor the comparison of subsets is well supported by state-of-the-art techniques. In this paper we present Domino, a novel multiform visualization technique for effectively representing subsets and the relationships between them. By providing comprehensive tools to arrange, combine, and extract subsets, Domino allows users to create both common visualization techniques and advanced visualizations tailored to specific use cases. In addition to the novel technique, we present an implementation that enables analysts to manage the wide range of options that our approach offers. Innovative interactive features such as placeholders and live previews support rapid creation of complex analysis setups. We introduce the technique and the implementation using a simple example and demonstrate scalability and effectiveness in a use case from the field of cancer genomics.


K. Grewen, M. Burchinal, C. Vachet, S. Gouttard, J.H. Gilmore, W. Lin, J. Johns, M. Elam, G. Gerig.
“Prenatal cocaine effects on brain structure in early infancy,” In NeuroImage, Vol. 101, pp. 114--123. November, 2014.
DOI: 10.1016/j.neuroimage.2014.06.070
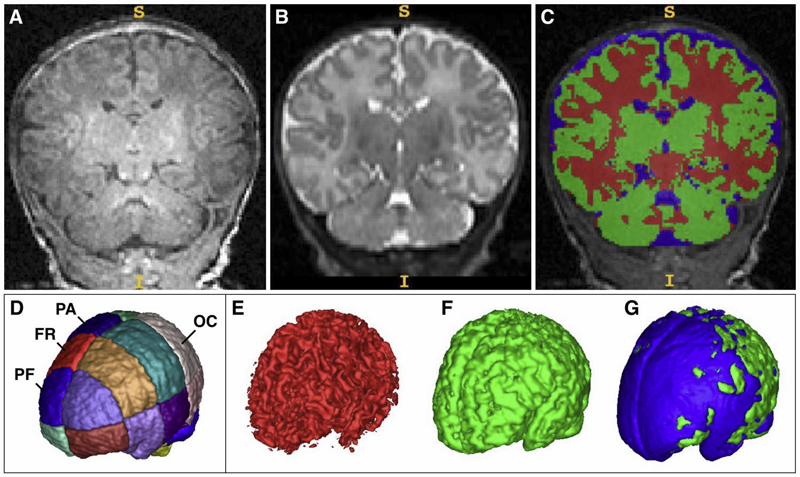
Prenatal cocaine exposure (PCE) is related to subtle deficits in cognitive and behavioral function in infancy, childhood and adolescence. Very little is known about the effects of in utero PCE on early brain development that may contribute to these impairments. The purpose of this study was to examine brain structural differences in infants with and without PCE. We conducted MRI scans of newborns (mean age = 5 weeks) to determine cocaine's impact on early brain structural development. Subjects were three groups of infants: 33 with PCE co-morbid with other drugs, 46 drug-free controls and 40 with prenatal exposure to other drugs (nicotine, alcohol, marijuana, opiates, SSRIs) but without cocaine. Infants with PCE exhibited lesser total gray matter (GM) volume and greater total cerebral spinal fluid (CSF) volume compared with controls and infants with non-cocaine drug exposure. Analysis of regional volumes revealed that whole brain GM differences were driven primarily by lesser GM in prefrontal and frontal brain regions in infants with PCE, while more posterior regions (parietal, occipital) did not differ across groups. Greater CSF volumes in PCE infants were present in prefrontal, frontal and parietal but not occipital regions. Greatest differences (GM reduction, CSF enlargement) in PCE infants were observed in dorsal prefrontal cortex. Results suggest that PCE is associated with structural deficits in neonatal cortical gray matter, specifically in prefrontal and frontal regions involved in executive function and inhibitory control. Longitudinal study is required to determine whether these early differences persist and contribute to deficits in cognitive functions and enhanced risk for drug abuse seen at school age and in later life.


C.E. Gritton.
“Ringing Instabilities in Particle Methods,” Note: M.S. in Computational Engineering and Science, advisor Martin Berzins, School of Computing, University of Utah, August, 2014.
Particle methods have been used in fields ranging from fluid dynamics to plasma physics. The Particle-In-Cell method and the family of methods that are an extension of it are a combination of both Lagrangian and Eularian methods. In this thesis, we present a brief survey of some of the methods and their key components. We show the different methods by which spatial derviates are computed. We propose a method of showing how the so-called "ringing instabilies" associated with particle methods arise and a means to remove them. We also propose that the underlying nodal scheme plays a key role in the stability of the method. Lastly, different particle methods are explored through numerical simulations and compared against an analytic solution.


Y. Gur, C.R. Johnson.
“Generalized HARDI Invariants by Method of Tensor Contraction,” In Proceedings of the 2014 IEEE International Symposium on Biomedical Imaging (ISBI), pp. 718--721. April, 2014.
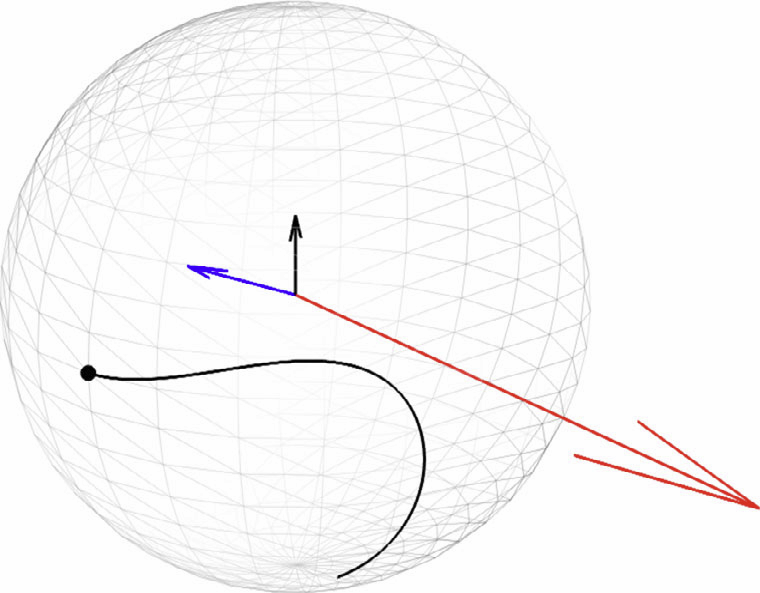
We propose a 3D object recognition technique to construct rotation invariant feature vectors for high angular resolution diffusion imaging (HARDI). This method uses the spherical harmonics (SH) expansion and is based on generating rank-1 contravariant tensors using the SH coefficients, and contracting them with covariant tensors to obtain invariants. The proposed technique enables the systematic construction of invariants for SH expansions of any order using simple mathematical operations. In addition, it allows construction of a large set of invariants, even for low order expansions, thus providing rich feature vectors for image analysis tasks such as classification and segmentation. In this paper, we use this technique to construct feature vectors for eighth-order fiber orientation distributions (FODs) reconstructed using constrained spherical deconvolution (CSD). Using simulated and in vivo brain data, we show that these invariants are robust to noise, enable voxel-wise classification, and capture meaningful information on the underlying white matter structure.
Keywords: Diffusion MRI, HARDI, invariants


C. Hamani, B.O. Amorim, A.L. Wheeler, M. Diwan, K. Driesslein, L. Covolan, C.R. Butson, J.N. Nobrega.
“Deep brain stimulation in rats: Different targets induce similar antidepressant-like effects but influence different circuits,” In Neurobiology of Disease, Vol. 71, Elsevier Inc., pp. 205--214. August, 2014.
ISSN: 1095-953X
DOI: 10.1016/j.nbd.2014.08.007
PubMed ID: 25131446
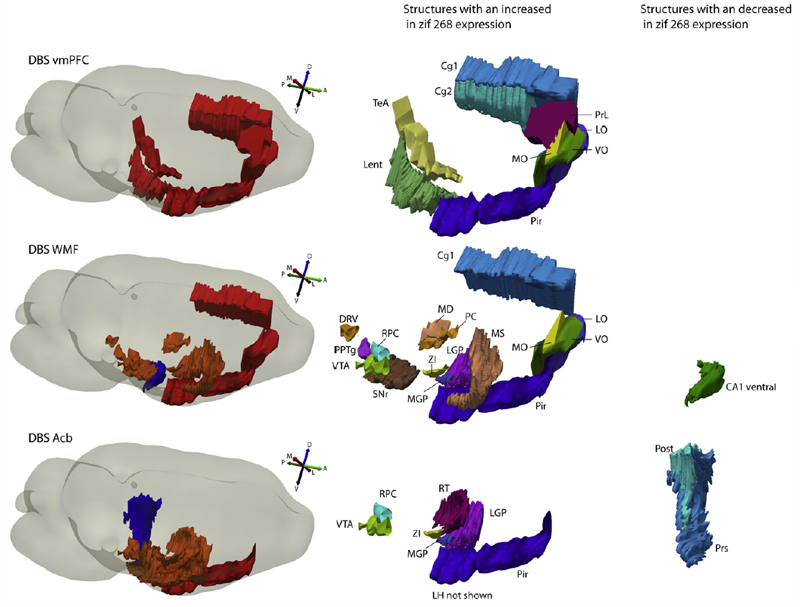
Recent studies in patients with treatment-resistant depression have shown similar results with the use of deep brain stimulation (DBS) in the subcallosal cingulate gyrus (SCG), ventral capsule/ventral striatum (VC/VS) and nucleus accumbens (Acb). As these brain regions are interconnected, one hypothesis is that by stimulating these targets one would just be influencing different relays in the same circuitry. We investigate behavioural, immediate early gene expression, and functional connectivity changes in rats given DBS in homologous regions, namely the ventromedial prefrontal cortex (vmPFC), white matter fibers of the frontal region (WMF) and nucleus accumbens. We found that DBS delivered to the vmPFC, Acb but not WMF induced significant antidepressant-like effects in the FST (31\%, 44\%, and 17\% reduction in immobility compared to controls). Despite these findings, stimulation applied to these three targets induced distinct patterns of regional activity and functional connectivity. While animals given vmPFC DBS had increased cortical zif268 expression, changes after Acb stimulation were primarily observed in subcortical structures. In animals receiving WMF DBS, both cortical and subcortical structures at a distance from the target were influenced by stimulation. In regards to functional connectivity, DBS in all targets decreased intercorrelations among cortical areas. This is in contrast to the clear differences observed in subcortical connectivity, which was reduced after vmPFC DBS but increased in rats receiving Acb or WMF stimulation. In conclusion, results from our study suggest that, despite similar antidepressant-like effects, stimulation of the vmPFC, WMF and Acb induce distinct changes in regional brain activity and functional connectivity.
Keywords: Anterior capsule, Connectivity, Deep brain stimulation, Depression, Nucleus accumbens, Prefrontal cortex

C.D. Hansen, M. Chen, C.R. Johnson, A.E. Kaufman, H. Hagen (Eds.).
“Scientific Visualization: Uncertainty, Multifield, Biomedical, and Scalable Visualization,” Mathematics and Visualization, Springer, 2014.
ISBN: 978-1-4471-6496-8


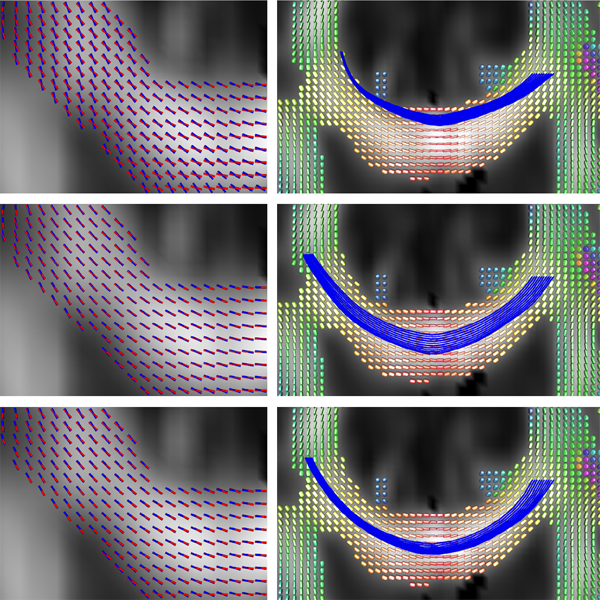
X. Hao, K. Zygmunt, R.T. Whitaker, P.T. Fletcher.
“Improved Segmentation of White Matter Tracts with Adaptive Riemannian Metrics,” In Medical Image Analysis, Vol. 18, No. 1, pp. 161--175. Jan, 2014.
DOI: 10.1016/j.media.2013.10.007
PubMed ID: 24211814
We present a novel geodesic approach to segmentation of white matter tracts from diffusion tensor imaging (DTI). Compared to deterministic and stochastic tractography, geodesic approaches treat the geometry of the brain white matter as a manifold, often using the inverse tensor field as a Riemannian metric. The white matter pathways are then inferred from the resulting geodesics, which have the desirable property that they tend to follow the main eigenvectors of the tensors, yet still have the flexibility to deviate from these directions when it results in lower costs. While this makes such methods more robust to noise, the choice of Riemannian metric in these methods is ad hoc. A serious drawback of current geodesic methods is that geodesics tend to deviate from the major eigenvectors in high-curvature areas in order to achieve the shortest path. In this paper we propose a method for learning an adaptive Riemannian metric from the DTI data, where the resulting geodesics more closely follow the principal eigenvector of the diffusion tensors even in high-curvature regions. We also develop a way to automatically segment the white matter tracts based on the computed geodesics. We show the robustness of our method on simulated data with different noise levels. We also compare our method with tractography methods and geodesic approaches using other Riemannian metrics and demonstrate that the proposed method results in improved geodesics and segmentations using both synthetic and real DTI data.
Keywords: Conformal factor, Diffusion tensor imaging, Front-propagation, Geodesic, Riemannian manifold


J. Hinkle, P.T. Fletcher, S. Joshi .
“Intrinsic Polynomials for Regression on Riemannian Manifolds,” In Journal of Mathematical Imaging and Vision, pp. 1-21. 2014.
We develop a framework for polynomial regression on Riemannian manifolds. Unlike recently developed spline models on Riemannian manifolds, Riemannian polynomials offer the ability to model parametric polynomials of all integer orders, odd and even. An intrinsic adjoint method is employed to compute variations of the matching functional, and polynomial regression is accomplished using a gradient-based optimization scheme. We apply our polynomial regression framework in the context of shape analysis in Kendall shape space as well as in diffeomorphic landmark space. Our algorithm is shown to be particularly convenient in Riemannian manifolds with additional symmetry, such as Lie groups and homogeneous spaces with right or left invariant metrics. As a particularly important example, we also apply polynomial regression to time-series imaging data using a right invariant Sobolev metric on the diffeomorphism group. The results show that Riemannian polynomials provide a practical model for parametric curve regression, while offering increased flexibility over geodesics.


T. Hollt, A. Magdy, P. Zhan, G. Chen, G. Gopalakrishnan, I. Hoteit, C.D. Hansen, M. Hadwiger.
“Ovis: A Framework for Visual Analysis of Ocean Forecast Ensembles,” In IEEE Transactions on Visualization and Computer Graphics (TVCG), Vol. PP, No. 99, pp. 1. 2014.
DOI: 10.1109/TVCG.2014.2307892
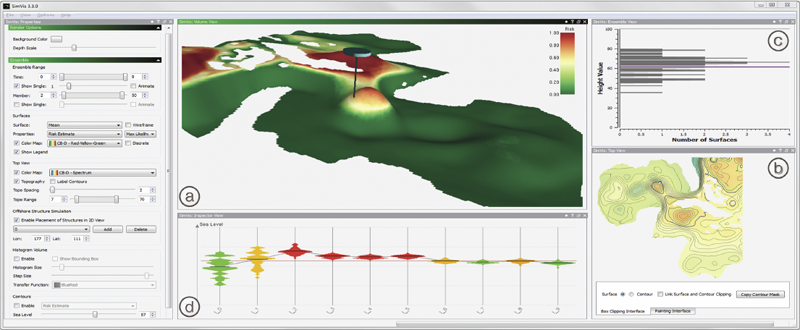
We present a novel integrated visualization system that enables interactive visual analysis of ensemble simulations of the sea surface height that is used in ocean forecasting. The position of eddies can be derived directly from the sea surface height and our visualization approach enables their interactive exploration and analysis. The behavior of eddies is important in different application settings of which we present two in this paper. First, we show an application for interactive planning of placement as well as operation of off-shore structures using real-world ensemble simulation data of the Gulf of Mexico. Off-shore structures, such as those used for oil exploration, are vulnerable to hazards caused by eddies, and the oil and gas industry relies on ocean forecasts for efficient operations. We enable analysis of the spatial domain, as well as the temporal evolution, for planning the placement and operation of structures. Eddies are also important for marine life. They transport water over large distances and with it also heat and other physical properties as well as biological organisms. In the second application we present the usefulness of our tool, which could be used for planning the paths of autonomous underwater vehicles, so called gliders, for marine scientists to study simulation data of the largely unexplored Red Sea.
Keywords: Ensemble Visualization, Ocean Visualization, Ocean Forecast, Risk Estimation


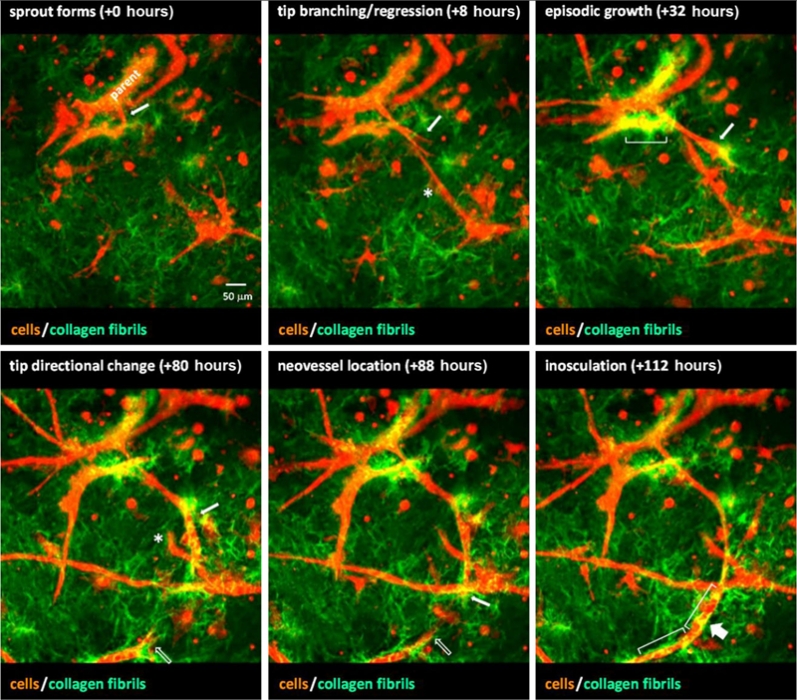
J.B. Hoying, U. Utzinger, J.A. Weiss.
“Formation of microvascular networks: role of stromal interactions directing angiogenic growth,” In Microcirculation, Vol. 21, No. 4, pp. 278--289. May, 2014.
DOI: 10.1111/micc.12115
PubMed ID: 24447042
PubMed Central ID: PMC4032604
In the adult, angiogenesis leads to an expanded microvascular network as new vessel segments are added to an existing microcirculation. Necessarily, growing neovessels must navigate through tissue stroma as they locate and grow toward other vessel elements. We have a growing body of evidence demonstrating that angiogenic neovessels reciprocally interact with the interstitial matrix of the stroma resulting in directed neovascular growth during angiogenesis. Given the compliance and the viscoelastic properties of collagen, neovessel guidance by the stroma is likely due to compressive strain transverse to the direction of primary tensile forces present during active tissue deformation. Similar stromal strains control the final network topology of the new microcirculation, including the distribution of arterioles, capillaries, and venules. In this case, stromal-derived stimuli must be present during the post-angiogenesis remodeling and maturation phases of neovascularization to have this effect. Interestingly, the preexisting organization of vessels prior to the start of angiogenesis has no lasting influence on the final, new network architecture. Combined, the evidence describes interplay between angiogenic neovessels and stroma that is important in directed neovessel growth and invasion. This dynamic is also likely a mechanism by which global tissue forces influence vascular form and function.
Keywords: angiogenesis, matrix, neovessel, remodeling, stroma


A. Humphrey, Q. Meng, M. Berzins, D. Caminha B.de Oliveira, Z. Rakamaric, G. Gopalakrishnan.
“Systematic Debugging Methods for Large-Scale HPC Computational Frameworks,” In Computing in Science Engineering, Vol. 16, No. 3, pp. 48--56. May, 2014.
ISSN: 1521-9615
DOI: 10.1109/MCSE.2014.11
Parallel computational frameworks for high performance computing (HPC) are central to the advancement of simulation based studies in science and engineering. Unfortunately, finding and fixing bugs in these frameworks can be extremely time consuming. Left unchecked, these bugs can drastically diminish the amount of new science that can be performed. This paper presents our systematic study of the Uintah Computational Framework, and our approaches to debug it more incisively. Our key insight is to leverage the modular structure of Uintah which lends itself to systematic debugging. In particular, we have developed a new approach based on Coalesced Stack Trace Graphs (CSTGs) that summarize the system behavior in terms of key control flows manifested through function invocation chains. We illustrate several scenarios how CSTGs could help efficiently localize bugs, and present a case study of how we found and fixed a real Uintah bug using CSTGs.
Keywords: Computational Modeling and Frameworks, Parallel Programming, Reliability, Debugging Aids


Y. Joon Ahn, C. Hoffmann, P. Rosen.
“Geometric constraints on quadratic Bézier curves using minimal length and energy,” In Journal of Computational and Applied Mathematics, Vol. 255, pp. 887--897. 2014.
This paper derives expressions for the arc length and the bending energy of quadratic Bézier curves. The formulas are in terms of the control point coordinates. For fixed start and end points of the Bézier curve, the locus of the middle control point is analyzed for curves of fixed arc length or bending energy. In the case of arc length this locus is convex. For bending energy it is not. Given a line or a circle and fixed end points, the locus of the middle control point is determined for those curves that are tangent to a given line or circle. For line tangency, this locus is a parallel line. In the case of the circle, the locus can be classified into one of six major types. In some of these cases, the locus contains circular arcs. These results are then used to implement fast algorithms that construct quadratic Bézier curves tangent to a given line or circle, with given end points, that minimize bending energy or arc length.


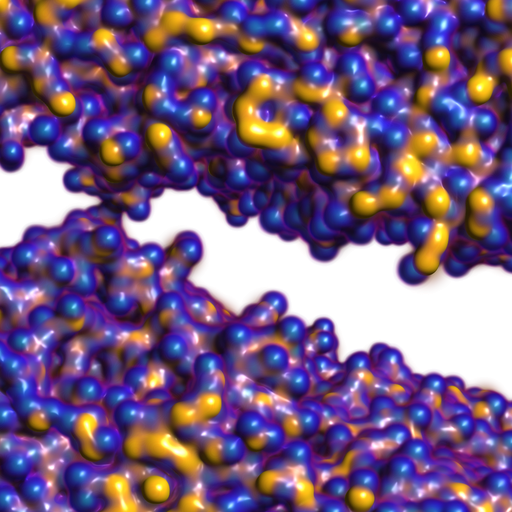
A. Knoll, I. Wald, P. Navratil, A. Bowen, K. Reda, M. E. Papka, K. Gaither.
“RBF Volume Ray Casting on Multicore and Manycore CPUs,” In Computer Graphics Forum, Vol. 33, No. 3, Edited by H. Carr and P. Rheingans and H. Schumann, Wiley-Blackwell, pp. 71--80. June, 2014.
DOI: 10.1111/cgf.12363
Modern supercomputers enable increasingly large N-body simulations using unstructured point data. The structures implied by these points can be reconstructed implicitly. Direct volume rendering of radial basis function (RBF) kernels in domain-space offers flexible classification and robust feature reconstruction, but achieving performant RBF volume rendering remains a challenge for existing methods on both CPUs and accelerators. In this paper, we present a fast CPU method for direct volume rendering of particle data with RBF kernels. We propose a novel two-pass algorithm: first sampling the RBF field using coherent bounding hierarchy traversal, then subsequently integrating samples along ray segments. Our approach performs interactively for a range of data sets from molecular dynamics and astrophysics up to 82 million particles. It does not rely on level of detail or subsampling, and offers better reconstruction quality than structured volume rendering of the same data, exhibiting comparable performance and requiring no additional preprocessing or memory footprint other than the BVH. Lastly, our technique enables multi-field, multi-material classification of particle data, providing better insight and analysis.


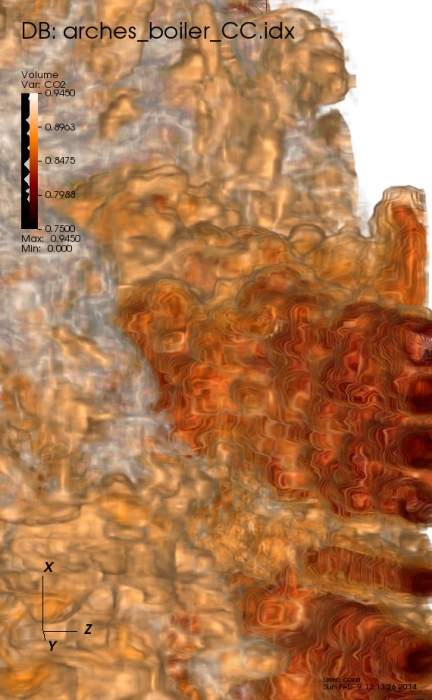
S. Kumar, C. Christensen, P.-T. Bremer, E. Brugger, V. Pascucci, J. Schmidt, M. Berzins, H. Kolla, J. Chen, V. Vishwanath, P. Carns, R. Grout.
“Fast Multi-Resolution Reads of Massive Simulation Datasets,” In Proceedings of the International Supercomputing Conference ISC'14, Leipzig, Germany, June, 2014.
Today's massively parallel simulation code can produce output ranging up to many terabytes of data. Utilizing this data to support scientific inquiry requires analysis and visualization, yet the sheer size of the data makes it cumbersome or impossible to read without computational resources similar to the original simulation. We identify two broad classes of problems for reading data and present effective solutions for both. The first class of data reads depends on user requirements and available resources. Tasks such as visualization and user-guided analysis may be accomplished using only a subset of variables with restricted spatial extents at a reduced resolution. The other class of reads require full resolution multi-variate data to be loaded, for example to restart a simulation. We show that utilizing the hierarchical multi-resolution IDX data format enables scalable and efficient serial and parallel read access on a variety of hardware from supercomputers down to portable devices. We demonstrate interactive view-dependent visualization and analysis of massive scientific datasets using low-power commodity hardware, and we compare read performance with other parallel file formats for both full and partial resolution data.


S. Kumar, J. Edwards, P.-T. Bremer, A. Knoll, C. Christensen, V. Vishwanath, P. Carns, J.A. Schmidt, V. Pascucci.
“Efficient I/O and storage of adaptive-resolution data,” In Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, IEEE Press, pp. 413--423. 2014.
DOI: 10.1109/SC.2014.39
We present an efficient, flexible, adaptive-resolution I/O framework that is suitable for both uniform and Adaptive Mesh Refinement (AMR) simulations. In an AMR setting, current solutions typically represent each resolution level as an independent grid which often results in inefficient storage and performance. Our technique coalesces domain data into a unified, multiresolution representation with fast, spatially aggregated I/O. Furthermore, our framework easily extends to importance-driven storage of uniform grids, for example, by storing regions of interest at full resolution and nonessential regions at lower resolution for visualization or analysis. Our framework, which is an extension of the PIDX framework, achieves state of the art disk usage and I/O performance regardless of resolution of the data, regions of interest, and the number of processes that generated the data. We demonstrate the scalability and efficiency of our framework using the Uintah and S3D large-scale combustion codes on the Mira and Edison supercomputers.


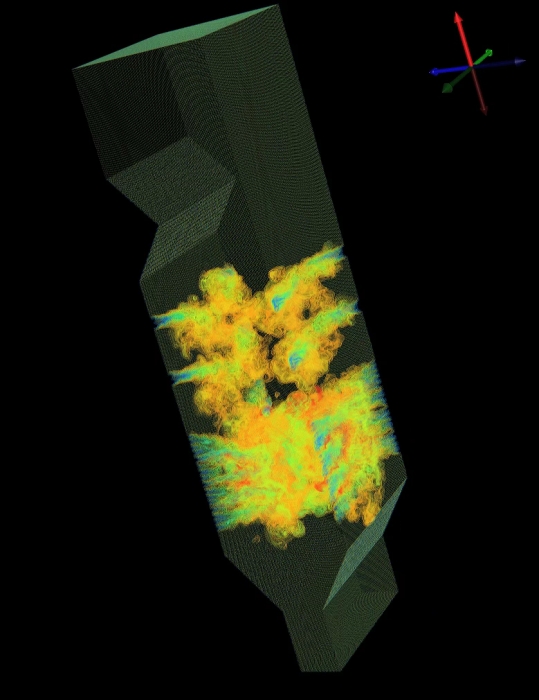
A.G. Landge, V. Pascucci, A. Gyulassy, J.C. Bennett, H. Kolla, J. Chen, P.-T. Bremer.
“In-situ feature extraction of large scale combustion simulations using segmented merge trees,” In Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis (SC 2014), New Orleans, Louisana, IEEE Press, Piscataway, NJ, USA pp. 1020--1031. 2014.
ISBN: 978-1-4799-5500-8
DOI: 10.1109/SC.2014.88
The ever increasing amount of data generated by scientific simulations coupled with system I/O constraints are fueling a need for in-situ analysis techniques. Of particular interest are approaches that produce reduced data representations while maintaining the ability to redefine, extract, and study features in a post-process to obtain scientific insights.
This paper presents two variants of in-situ feature extraction techniques using segmented merge trees, which encode a wide range of threshold based features. The first approach is a fast, low communication cost technique that generates an exact solution but has limited scalability. The second is a scalable, local approximation that nevertheless is guaranteed to correctly extract all features up to a predefined size. We demonstrate both variants using some of the largest combustion simulations available on leadership class supercomputers. Our approach allows state-of-the-art, feature-based analysis to be performed in-situ at significantly higher frequency than currently possible and with negligible impact on the overall simulation runtime.


J.D. Lewis, A.C. Evans, J.R. Pruett, K. Botteron, L. Zwaigenbaum, A. Estes, G. Gerig, L. Collins, P. Kostopoulos, R. McKinstry, S. Dager, S. Paterson, R. Schultz, M. Styner, H. Hazlett, J. Piven, IBIS network.
“Network inefficiencies in autism spectrum disorder at 24 months,” In Translational Psychiatry, Vol. 4, No. 5, Nature Publishing Group, pp. e388. May, 2014.
DOI: 10.1038/tp.2014.24
Autism Spectrum Disorder (ASD) is a developmental disorder defined by behavioural symptoms that emerge during the first years of life. Associated with these symptoms are differences in the structure of a wide array of brain regions, and in the connectivity between these regions. However, the use of cohorts with large age variability and participants past the generally recognized age of onset of the defining behaviours means that many of the reported abnormalities may be a result of cascade effects of developmentally earlier deviations. This study assessed differences in connectivity in ASD at the age at which the defining behaviours first become clear. The participants were 113 24-month-olds at high risk for ASD, 31 of whom were classified as ASD, and 23 typically developing 24-month-olds at low risk for ASD. Utilizing diffusion data to obtain measures of the length and strength of connections between anatomical regions, we performed an analysis of network efficiency. Our results showed significantly decreased local and global efficiency over temporal, parietal, and occipital lobes in high-risk infants classified as ASD, relative to both low- and high-risk infants not classified as ASD. The frontal lobes showed only a reduction in global efficiency in Broca's area. Additionally, these same regions showed an inverse relation between efficiency and symptom severity across the high-risk infants. The results suggest delay or deficits in infants with ASD in the optimization of both local and global aspects of network structure in regions involved in processing auditory and visual stimuli, language, and nonlinguistic social stimuli.
Keywords: autism, infant siblings, connectivity, network analysis, efficiency


A. Lex, N. Gehlenborg, H. Strobelt, R. Vuillemot,, H. Pfister.
“UpSet: Visualization of Intersecting Sets,” In IEEE Transactions on Visualization and Computer Graphics (InfoVis '14), Vol. 20, No. 12, pp. 1983--1992. 2014.
ISSN: 1077-2626
Understanding relationships between sets is an important analysis task that has received widespread attention in the visualization community. The major challenge in this context is the combinatorial explosion of the number of set intersections if the number of sets exceeds a trivial threshold. In this paper we introduce UpSet, a novel visualization technique for the quantitative analysis of sets, their intersections, and aggregates of intersections. UpSet is focused on creating task-driven aggregates, communicating the size and properties of aggregates and intersections, and a duality between the visualization of the elements in a dataset and their set membership. UpSet visualizes set intersections in a matrix layout and introduces aggregates based on groupings and queries. The matrix layout enables the effective representation of associated data, such as the number of elements in the aggregates and intersections, as well as additional summary statistics derived from subset or element attributes. Sorting according to various measures enables a task-driven analysis of relevant intersections and aggregates. The elements represented in the sets and their associated attributes are visualized in a separate view. Queries based on containment in specific intersections, aggregates or driven by attribute filters are propagated between both views. We also introduce several advanced visual encodings and interaction methods to overcome the problems of varying scales and to address scalability. UpSet is web-based and open source. We demonstrate its general utility in multiple use cases from various domains.


W. Liu, S.P. Awate, J.S. Anderson, P.T. Fletcher.
“A functional network estimation method of resting-state fMRI using a hierarchical Markov random field,” In NeuroImage, Vol. 100, pp. 520--534. 2014.
ISSN: 1053-8119
DOI: 10.1016/j.neuroimage.2014.06.001
We propose a hierarchical Markov random field model for estimating both group and subject functional networks simultaneously. The model takes into account the within-subject spatial coherence as well as the between-subject consistency of the network label maps. The statistical dependency between group and subject networks acts as a regularization, which helps the network estimation on both layers. We use Gibbs sampling to approximate the posterior density of the network labels and Monte Carlo expectation maximization to estimate the model parameters. We compare our method with two alternative segmentation methods based on K-Means and normalized cuts, using synthetic and real data. The experimental results show that our proposed model is able to identify both group and subject functional networks with higher accuracy on synthetic data, more robustness, and inter-session consistency on the real data.
Keywords: Resting-state functional MRI, Segmentation, Functional connectivity, Hierarchical Markov random field, Bayesian


Shusen Liu, Bei Wang, J.J. Thiagarajan, P.-T. Bremer, V. Pascucci.
“Visual Exploration of High-Dimensional Data: Subspace Analysis through Dynamic Projections,” SCI Technical Report, No. UUSCI-2014-003, SCI Institute, University of Utah, 2014.
Understanding high-dimensional data is rapidly becoming a central challenge in many areas of science and engineering. Most current techniques either rely on manifold learning based techniques which typically create a single embedding of the data or on subspace selection to find subsets of the original attributes that highlight the structure. However, the former creates a single, difficult-to-interpret view and assumes the data to be drawn from a single manifold, while the latter is limited to axis-aligned projections with restrictive viewing angles. Instead, we introduce ideas based on subspace clustering that can faithfully represent more complex data than the axis-aligned projections, yet do not assume the data to lie on a single manifold. In particular, subspace clustering assumes that the data can be represented by a union of low-dimensional subspaces, which can subsequently be used for analysis and visualization. In this paper, we introduce new techniques to reliably estimate both the intrinsic dimension and the linear basis of a mixture of subspaces extracted through subspace clustering. We show that the resulting bases represent the high-dimensional structures more reliably than traditional approaches. Subsequently, we use the bases to define different “viewpoints”, i.e., different projections onto pairs of basis vectors, from which to visualize the data. While more intuitive than non-linear projections, interpreting linear subspaces in terms of the original dimensions can still be challenging. To address this problem, we present new, animated transitions between different views to help the user navigate and explore the high-dimensional space. More specifically, we introduce the view transition graph which contains nodes for each subspace viewpoint and edges for potential transition between views. The transition graph enables users to explore both the structure within a subspace and the relations between different subspaces, for better understanding of the data. Using a number of case studies on well-know reference datasets, we demonstrate that the interactive exploration through such dynamic projections provides additional insights not readily available from existing tools.
Keywords: High-dimensional data, Subspace, Dynamic projection